Hi,
I have got these results from tophat-fusion:
chr15-chr17 74317267 38506035 ff 1 13 1 7 43 33 10.000000 @ 13 26 38 51 63 @ CCAGCCCAGAGGCTGCCAGCACTCCCAGGGACCCTATTGACGTTGACCTG GTGAGATGGGTTTGAGGTCTGAAGGGGTGGTGGT
GGGGCCCAGCAGGTCA @ TGCTCCCACTGTGGGTGTGGACAACCTGACTCCCTCCCCTCCATACCCAG GGCTTCTTCCGCCGCAGCATCCAGAAGAACATGGTGTACACGTGTCACCG @ 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 0 0 0 0
0 0 0 @ 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 @ -8418:28 -8481:-1326 -8394:-1415 -8411:-1464 -8421:-1464 -8479:-1417 -8461:-1462 -8476:-1471 -8794:-5041 -872
9:-5209 -8803:-5220 -8834:-5250 -8799:-5285
chr15-chr17 74325754 38504567 ff 5 13 9 1 54 39 15.480002 @ 14 30 41 53 68 @ GGTCTTCCTGCCCAACAGCAACCACGTGGCCAGTGGCGCCGGGGAGGCAG GTAGGGAGGTGGGTAGGGCAGTGGCCTGGGTGCT
GCCCGCCAGGGTCTGG @ AAGGATGGCGACCTAGGTCTCTAACTGCCCCTCCCCTCTTCTCTCTCTAG CCATTGAGACCCAGAGCAGCAGTTCTGAAGAGATAGTGCCCAGCCCTCCC @ 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 4 4 4 4 4 4 4 4 4 4
4 4 3 @ 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 3 2 2 2 2 1 1 1 1 1 1 1 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 @ 11:-3 26:6 8:51 66:4 76:4 93:53 6:142 69:1496 -307:-3573 -242:-3741 -316:-3752 -347:-3782 -312:-3817
chr15-chr17 74326139 38500724 ff 8 13 10 3 63 60 1.859375 @ 15 27 41 55 67 @ GCATACCATAACAGGAAACTGGACTCCCCATGACCTTAACTCTGCTCTCT CAGTCTACACCCATCATTTCCCCAAGTGTTTCTG
CAAAGGCCACCTACCT @ GGGCTGCCAGAGTCACCCCTTCCACTGCCTTGGCCACCTTCTCCAGAGGG CTGGAGAGAAGCTGGGATCTGAGACCTTGGTCTCCAGCCCCTGTCTCTTC @ 8 8 8 8 8 8 8 8 8 8 8 8 8 8 8 8 7 7 7 7 7 6 6 6 6 6 6 6 6 6 6 6 6 5 5 5 5 4 4 4 4 4 4 4 4 4 4
4 4 4 @ 8 8 8 8 8 8 8 8 8 8 8 8 8 5 5 5 5 5 5 5 5 5 5 5 5 4 4 4 4 4 4 4 4 4 4 4 4 4 4 3 3 3 3 2 2 2 2 2 2 2 @ 38:61 73:26 69:91 143:102 78:270 396:3840 411:3849 393:3894 451:3847 461:3847 478:3896 391:3985 454:5339
chr15-chr17 74326818 38487647 rr 15 4 18 1 8515 61 0.675556 @ 13 21 35 46 57 @ GTTTTCGGCATCTGAGTCTTCCGAGCTGCTGATCACCACAACGCGTTCCT CTGAAACGGGGAAGGGGAGATGCATTATGAACTG
CCTAGGATGCATGCTG @ TGCCACCCTCCACAGTCCCTTCCCACCCCAGCCCCCACACCCAGACTTAC TGGCTGGGGATGGTGTGCTATATCCACTAACTGGAAGCTGGTGCTGGAGA @ 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 12 12 12 12 12 12 12 10 10 10 10 10 10 10 9 9 9
9 8 8 6 6 6 6 6 6 6 6 6 5 5 5 5 5 4 @ 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 15 14 14 14 14 13 13 12 11 10 10 10 10 10 9 9 9 9 9 9 9 9 9 7 7 6 6 6 6 5 5 5 @ 23:8 44:21 8638:4 8622:44
chr15-chr17 74325591 38504567 ff 2 1 2 26 34 47 10.500000 @ 15 28 44 56 66 @ CCCAGTGCCCCAGGAAGGTCATCAAGATGGAGTCTGAGGAGGGGAAGGAG GCAAGGTTGGCTCGGAGCTCCCCGGAGCAGCCCA
GGCCCAGCACCTCCAA @ AAGGATGGCGACCTAGGTCTCTAACTGCCCCTCCCCTCTTCTCTCTCTAG CCATTGAGACCCAGAGCAGCAGTTCTGAAGAGATAGTGCCCAGCCCTCCC @ 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 @ 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 1 1 1 1 1 0 0 0 @ -106:4
I have the feeling that the number of reads spanning the fusion is too low and predictions might not be accurate, but, on the other hand we were expecting these fusions but not with low number of reads. More over the the number of contradicting reads are also high.
I am really confused, what shall I do? can I take these results and confirm that fusion takes place or it is just by chance that I am seeing this.
Please help with your expert opinion and how you would have judged this result.
Thank you

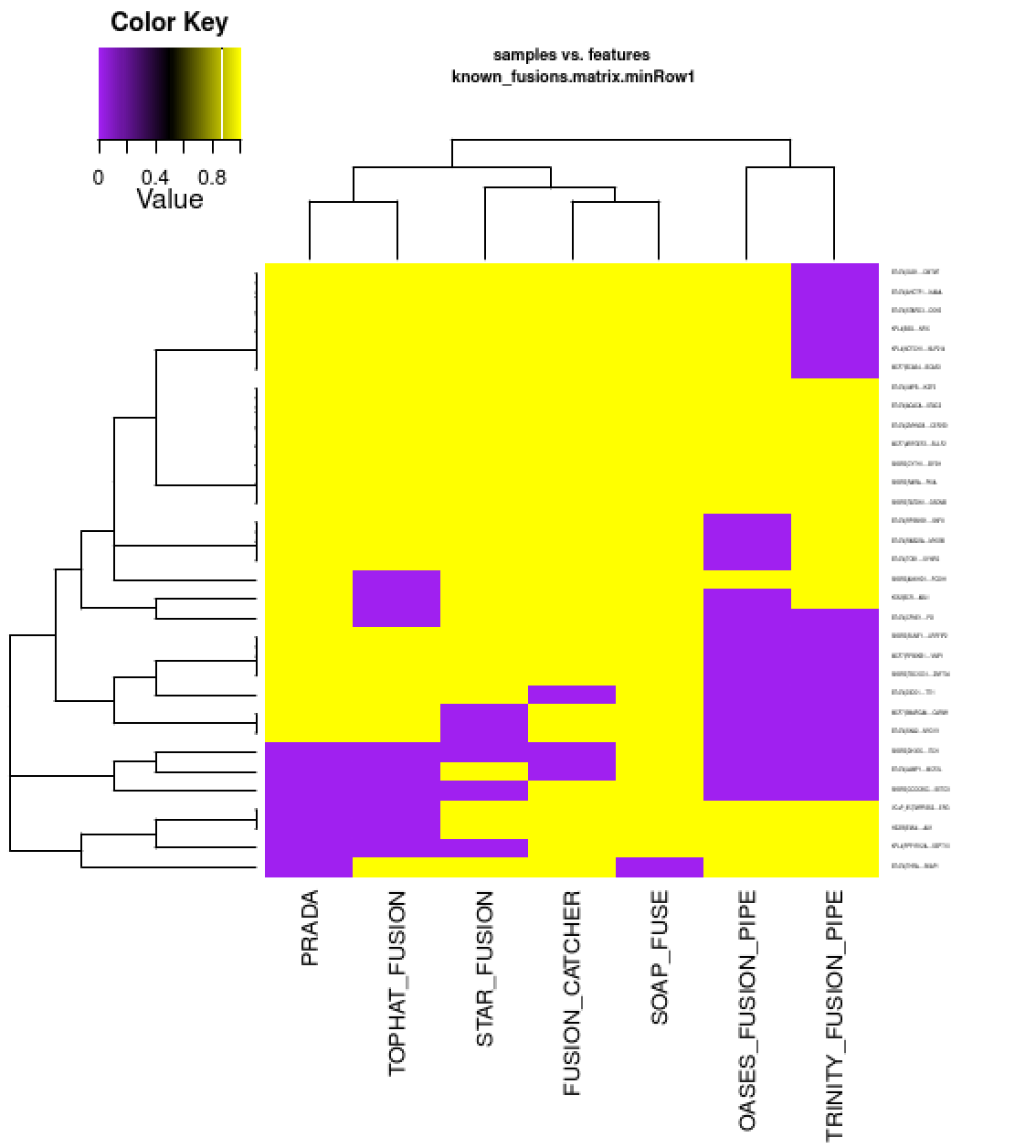

I have a feeling from my analysis that TopHat fusion gives so many fusions and lot of false positive results, whereas STAR is better , you can even have trust on MapSplice2.