I am analyzing 20 Agilent microarrays from humans (20 subjects, two conditions) and yet there is no differentially expressed gene after adjusting for multiple testing (BH). I used Limma in R, tried different background correction as well as normalization methods (within and between arrays), I also tried to remove the least expressed probes to try to boost the signal. All the array quality checks looked fine.
The question is, if there are no significantly differentially expressed genes, what can be done? Has anyone ever tried to get some results out of this situation or has a good paper/link to suggest?
Thanks all!

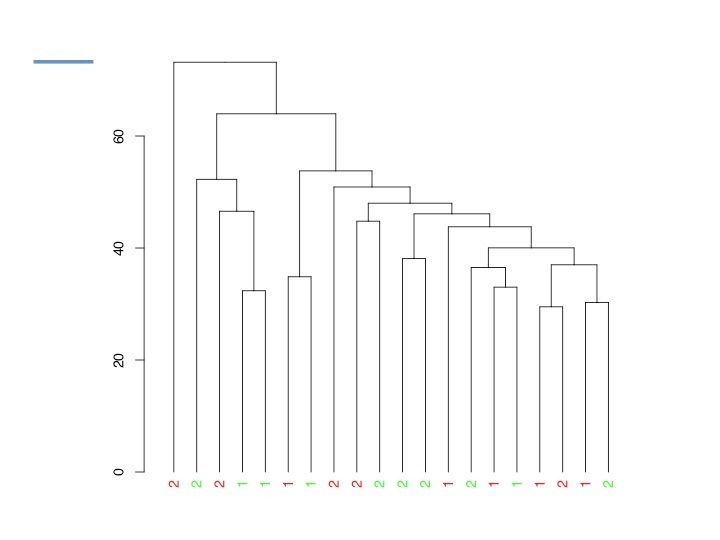

How is the correlation between biological replicates ?
There are no biological replicates, let's say I have condition A on 10 subjects and condition B on the other 10
you can always look at the fold change values.
but they need to be significant, no?
Variance would be very high within the groups thats why you do not get significant DEGs between the groups, so better to try out as I suggested in the answer below,
lets see what happens then.