I have ChIP-seq data with peaks called by MACS in peaks.bed and summits.bed files. From here I would like to visualise the peaks on either the IGV or UCSC' Genome Browser. I understand that the bed files give me the positions and size of the peaks called by MACS, but how do I actually visualise the height / distribution of the peaks? Do I have to get the genome coverage for each base from my bam file, then filter for peak regions listed in the bed files, and then convert to a bedgraph, wig, or bigwig format?
1. Use -w and -S parameters, which gives you two 'wig' files: One for Control and another for TREATMENT. Wiggle (wig) file gives you the distribution and coverage (height) of peaks which you can upload on UCSC or IGV . Read more about the other parameters here.
---Example:
macs14 -c <CONTROL_FILE> -t <TREATMENT_FILE> -n OUTPUT_NAME -f BAM -g hs -w -S
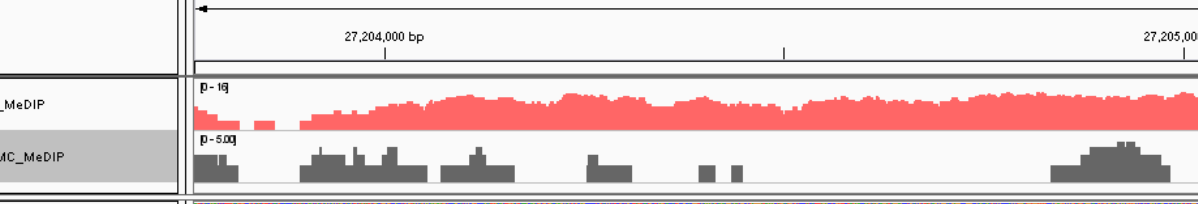
2. If you want to just show the coverage over predicted peak regions by MACS, use BEDtools to extract only the portion of 'wig' file overlapping with peak regions (BED file).
I would recommend using MACS2. I recently added a method for installing this virtualenv - running Python based software in isolation.
Using the -B flag you will get a two bedGraph files (one for treatments and one for the lambda model based on the control). bedGraph files are much, much smaller than WIG files. You can also use the --SPMR flag to normalise the coverage based on millions of reads used by MACS, which is very useful when comparing different experiments.
If you have a web server you can view the coverage plots on UCSC. But you need to convert bedGraph to bigWig first. The programs below can be downloaded from http://hgdownload.soe.ucsc.edu/admin/exe/linux.x86_64/.
# generate chromosome lengths for genome
fetchChromSizes.sh mm9 > mm9.chrom.sizes
# clip the ends of reads that overlap the ends of chromosomes
bedClip treat.bedgraph mm9.chrom.sizes treat.clipped
# convert clipped bedGraph to bigWig
bedGraphToBigWig treat.clipped mm9.chrom.sizes treat.bw
However, you can directly view bedGraph files in IGB and possibly also IGV (not checked recently), without conversion or need of a web server.
Looking at thousands of peaks on a genome browser is time consuming and usually not very productive. What I normally do is visualise the peaks on a genome browser as a quality control and then use summary plots like heatmaps or meta profiles either around the called peaks or using some other feature like gene start.
I recommend you to look at the deepTools package that handles all processing and visualisation steps. Here is a video I made with a quick example.
Use of this site constitutes acceptance of our User Agreement and Privacy Policy.


Thanks for the great reply, could you provide an example for extracting the overlapping regions?